Cloud-Based eDNA Data Management & Exploration Platform
The Amplicon Sequence Variant (ASV) Registry is a web application developed for the Smithsonian Institution’s Ocean DNA initiative and Jonah Ventures’ marine eDNA monitoring programs. It serves as both a centralized data registry and an interactive exploration platform for amplicon sequence variant data across multiple ocean biodiversity projects.
The system integrates sequence data, sample metadata, taxonomic assignments, and spatial context into a unified interface designed for cross-project comparison and standardized biodiversity reporting.
The Problem
Environmental DNA monitoring produces large volumes of sequence data that must be:
- Linked to detailed sample metadata
- Assigned taxonomy across evolving reference libraries
- Compared across projects with different workflows
- Exported in standardized formats for biodiversity databases such as GBIF
Researchers often rely on fragmented scripts or static outputs, making it difficult to investigate poorly matched sequences, compare taxa across projects, or maintain versioned taxonomy over time.
A scalable, structured platform was needed to manage ASV data across ocean monitoring efforts while supporting both exploration and downstream reporting.
The Solution
I developed a modular Shiny application structured around two core components: data registry and interactive exploration.
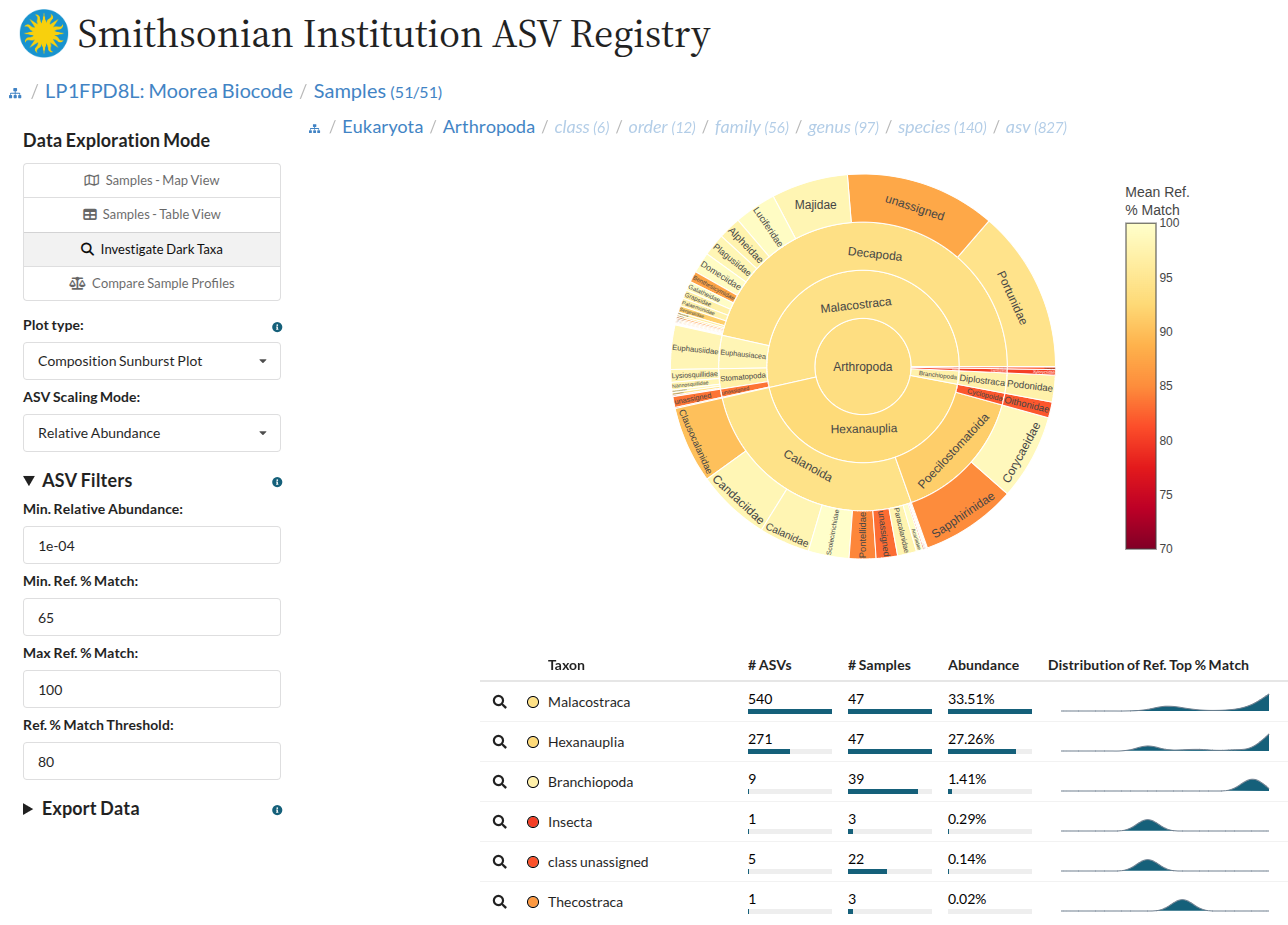
Dark Taxa Investigation:
- Interactive sunburst plots visualizing taxonomic composition with match-quality overlays
- Scatter plots highlighting poorly matched sequences by abundance and similarity
- ASV selection tools for exporting sequences for external analysis
Core Capabilities:
- Integration with GEOME via API for automated sample metadata retrieval
- AWS Lambda-based automated taxonomic assignment
- Interactive Leaflet maps with drawing tools for spatial subsetting
- Species accumulation curves, shared taxa analysis, and ordination plots
- Darwin Core-compliant exports for GBIF integration
- Taxonomy versioning support for temporal comparisons
- Flexible cross-project filtering
The result is a system that supports both exploratory analysis and standardized biodiversity data exchange.
Technical Architecture
The application was built using the golem framework to ensure modularity and long-term maintainability.
Key architectural components include:
- SQLite database backend for structured ASV and metadata storage
- Amazon S3 for spatial and project data assets
- AWS Lambda for asynchronous taxonomic assignment workflows
- Docker containerization with a two-stage build process
- ShinyProxy deployment on AWS EC2, providing session-level container isolation
- AWS Cognito authentication for secure user access
ShinyProxy ensures each user session runs in an isolated container instance, maintaining both security and reproducibility. The schema was designed with migration to production-scale database infrastructure in mind.
Outcome
What began as a prototype for exploring eDNA visualization approaches evolved into a production system managing data from multiple marine monitoring projects.
The ASV Registry currently supports cross-project biodiversity analysis and is positioned for expanded deployment within Smithsonian computational infrastructure, including integration with GBIF via standardized Darwin Core exports.
The project demonstrates how biodiversity informatics systems can be engineered as durable, modular platforms rather than ad hoc analysis scripts — bridging ecological genomics and production infrastructure.